PCR
PCR (Polymer Chain Reaction)
upscales minute concentrations of polynucletides to detectable
amounts. The fragments then are separated by electrophoresis with
respect to chainlength.
Scanned frationation patterns are highly superresolvable, since
the kernel is a somooth and perfectly reproducible parameter,
especially with modern microtechniques.
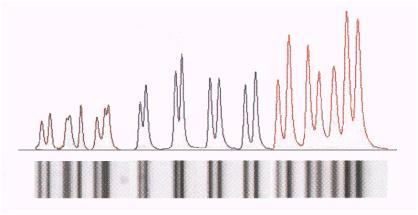
The table describes the process of
digital superresolution of PCR-data:
| Step |
Black
line: Original
data
Grey
solid: Superresolution
Green
line: Kernel |
| Perform the separation and scan
the pattern Modern
markers allow for exact mass-propotional detection of
polynucleotide patterns, just like in classical HPLC.
Import of the scanned data
into a resolver software is all, that's needed to achieve
approximate duplicate resolution and removal of peak
tails and asymmetries.
|
 |
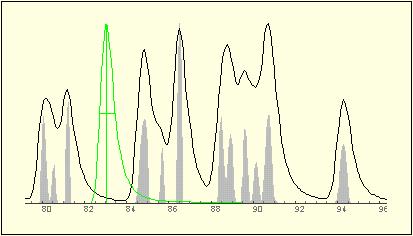
| Superresolve The example shows, how seemingly 8
peaks decompose into twelve ones.
Calculation time is less
than 5 seconds and can be fully automatted.
|
 |
Reducing sampling intervals,
High throughput optimization |
Please
consult the page 'Reducing
sampling times in chromatography'' |
Superresolution
requires robust baseline correction.
In PROANALYSI::PEAKS, you find baseline-routines
optimized especially for PCR.
The complete sequence of data analysis can be automatted and
perfomed by simply pressing a button.

